How-to’s
This section will explain how to make the most of pygamma-agreement’s customization possibilities.
Setting up your own positional dissimilarity
The only positionnal dissimilarity available in pygamma-agreement is the Positional Sporadic dissimilarity,
introduced by [mathet2015]. In this part, we’ll detail how to use your own for computing the gamma agreement.
A Dissimilarity class is a class that holds the information needed to compile a dissimilarity function.
It is made this way to optimize as much as possible the very costly computation of gamma. Let’s start by
inheriting the AbstractDissimilarity class :
from pygamma_agreement import AbstractDissimilarity, Unit
import numpy as np
from typing import Callable
class MyPositionalDissimilarity(AbstractDissimilarity):
def __init__(self, delta_empty=1.0):
super().__init__(delta_empty=delta_empty)
# Abstract methods overrides
def compile_d_mat(self) -> Callable[[np.ndarray, np.ndarray], float]:
...
def d(self, unit1: Unit, unit2: Unit) -> float:
...
This is essentially the minimum skeleton needed to define a new positional dissimilarity. You can also use additionnal attributes to make your dissimilarity more parametrizable.
For this example, the dissimilarity we’ll be implementing will be defined as such : for an integer \(p\),
thus, let’s redefine the constructor accordingly :
def __init__(self, p: int, delta_empty=1.0):
self.p = p
assert p > 0
super().__init__(delta_empty=delta_empty)
Great. Now, let’s explain what the d method is.
The AbstractDissimilarity.d method is essentially the python method that returns the dissimilarity between the two
non-empty given units. When the units are empty, any algorithm from pygamma-agreement that uses a dissimilarity
object will do the checking out of this method and use the delta_empty attribute accordingly.
So, you only have to implement the first part of \(d_p\)’s definition.
def d(self, unit1: Unit, unit2: Unit) -> float:
return ( abs(unit1.segment.start - unit2.segment.start)**self.p
+ abs(unit1.segment.end - unit2.segment.end)**self.p ) ** (1/self.p)
This method is essentially used in the gamma-cat computation, since it is too slow for the costly gamma algorithm.
It’s time to explain the essential part of the dissimilarity : the compile_d_mat method.
To minimize computation time of the gamma-agreement, the dissimilarities are used in a C-compiled form
generated by the numba library. To accomplish this, they need to keep the same signature, and to
only use native python or numpy types/operations internally.
Let’s write the MyPositionalDissimilarity.compile_d_mat method to better explain it :
def compile_d_mat(self) -> Callable[[np.ndarray, np.ndarray], float]:
# Calling self inside d_mat makes the compiler choke, so you need to copy attributes in locals.
p = self.p
delta_empty = self.delta_empty
from pygamma_agreement import dissimilarity_dec
@dissimilarity_dec # This decorator specifies that this function will be compiled.
def d_mat(unit1: np.ndarray, unit2: np.ndarray) -> float:
# We're in numba environment here, which means that only python/numpy types and operations will work.
return (abs(unit1[0] - unit2[0])**p
+ abs(unit1[1] - unit2[1])**p)**(1 / p) * delta_empty
return d_mat
You’ll notice that the units’ attributes are accessed by index. The correspondance is the following :
unit_array: np.ndarray
unit_object: Unit
unit_array[0] == unit_object.segment.start
unit_array[1] == unit_object.segment.end
unit_array[2] == unit_object.segment.end - unit_object.segment.start
Now, the dissimilarity is ready to be used !
from pygamma_agreement import Continuum
continuum: Continuum
dissim = MyPositionalDissimilarity(p=2, delta_empty=1.0)
gamma_results = continuum.compute_gamma(dissim)
Warning
A very important thing to note is that the structure of dissimilarities is not really compatible with changing attributes, because of the class structure and of compilation. It is advised do redefine your dissimilarities if you want to change attributes.
dissim.p = 3 # DON'T do that !
dissim = MyPositionalDissimilarity(p=3, delta_empty=1.0) # Redefine it instead.
Setting up your own categorical dissimilarity
For many reasons, string types are not easy to manipulate in numba njit’ed code.
Instead, category-to-category dissimilarities are pre-computed at the python level. Thus, there is a very simple
interface avaible : You just need to inherit the LambdaCategoricalDissimilarity, and override the
cat_dissim_func static method :
class MyCategoricalDissimilarity(LambdaCategoricalDissimilarity):
# Precomputation requires the category labels to be saved. Don't use this dissimilarity with
# a continuum containing unspecified categories
def __init__(self, labels: Iterable[str], delta_empty: float = 1.0):
super().__init__(labels, delta_empty)
@staticmethod
def cat_dissim_func(str1: str, str2: str) -> float:
return ... # Your categorical dissimilarity function. Results should be in [0, 1]
Beware that in reality, the resulting dissimilarity between categories a and b will be
cat_dissim_func(a, b) * delta_empty
Your new categorical dissimilarity is now ready. You can, for instance, use it in a combined categorical dissimilarity :
from pygamma_agreement import CombinedCategoricalDissimilarity, Continuum
continuum: Continuum
dissim = CombinedCategoricalDissimilarity(alpha=3, beta=1,
cat_dissim=MyCategoricalDissimilarity(continuum.categories))
gamma_results = continuum.compute_gamma(dissim)
Combining dissimilarities
The only combined dissimilarity (a dissmilarity that considers both positionning and categorizing of units)
natively available in pygamma-agreement is the one introduced by [mathet2015]
(the CombinedCategoricalDissimilarity):
Imagine instead that you want to adapt this this dissimilarity geometrically :
Let’s start by using the same skeleton as for a simple positional dissimilarity :
from pygamma_agreement import AbstractDissimilarity, Unit
import numpy as np
from typing import Callable
class MyCombinedDissimilarity(AbstractDissimilarity):
def __init__(self, alpha: float, beta: float,
pos_dissim: AbstractDissimilarity,
cat_dissim: CategoricalDissimilarity,
delta_empty=1.0):
self.alpha, self.beta = alpha, beta
self.pos_dissim, self.cat_dissim = pos_dissim, cat_dissim
super().__init__(cat_dissim.categories, delta_empty=delta_empty)
# Abstract methods overrides
def compile_d_mat(self) -> Callable[[np.ndarray, np.ndarray], float]:
...
def d(self, unit1: Unit, unit2: Unit) -> float:
...
Note
One important thing to note that if your dissimilarity takes categories into account, you must specify a set of categories to the super constructor. Here in this example, the cat_dissim part does take it into account, so its categories can be obtained directly.
The d method can simply make use of the other dissimilarities’ d s :
def d(self, unit1: Unit, unit2: Unit) -> float:
return ( self.pos_dissim.d(unit1, unit2)**(self.alpha)
* self.cat_dissim.d(unit1, unit2)**(self.beta) )
Moreover, you can access a dissimilarity’s numba-compiled function from the d_mat attribute, which is
usable in numba-compiled environment. Thus, compiling the dissimilarity function is pretty simple, and very
similar to a simple dissimilarity with only arithmetic operations. Let’s illustrate this :
def compile_d_mat(self) -> Callable[[np.ndarray, np.ndarray], float]:
alpha, beta = self.alpha, self.beta
pos, cat = self.pos_dissim.d_mat, self.cat_dissim.d_mat
# d_mat attribute contains the numba-compiled function
from pygamma_agreement import dissimilarity_dec
@dissimilarity_dec
def d_mat(unit1: np.ndarray, unit2: np.ndarray) -> float:
return pos(unit1, unit2)**alpha * cat(unit1, unit2)**beta
return d_mat
Then, there’s only the d method left to code.
def d(self, unit1: Unit, unit2: Unit):
return (self.pos_dissim.d(unit1, unit2)**self.alpha *
self.cat_dissim.d(unit1, unit2)**self.beta)
And that’s it ! Now, in theory, you have all the tools you need to compute the gamma-agreement with any dissimilarity.
continuum: Continuum
dissim = MyCombinedDissimilarity(alpha=3, beta=2,
pos_dissim=MyPositionalDissimilarity(),
cat_dissim=MyCategoricalDissimilarity(continuum.categories))
gamma_results = continuum.compute_gamma(dissim)
Generating random continua for comparison using the Statistical Sampler and the Corpus Shuffling Tool
If you want to measure your dissimilarity’s influence on the gamma-agreement depending on
certain possible errors between annotations, pygamma-agreement contains all the the needed tools :
the StatisticalContinuumSampler, which generates totally random continuua, and the
CorpusShufflingTool, that simulates errors when annotating a resource.
First, let’s generate a random reference for the corpus shuffling tool, which will act as the perfectly accurate annotation:
from pygamma_agreement import StatisticalContinuumSampler, CorpusShufflingTool, Continuum
sampler = StatisticalContinuumSampler()
# avg stands for average, std stands for standart deviation. All values are generated using normal distribution.
sampler.init_sampling_custom(["annotator_ref"],
avg_num_units_per_annotator=50, std_num_units_per_annotator=10,
avg_duration=15, std_duration=3,
avg_gap=5, std_gap=1,
categories=["Speaker1", "Speaker2", "Speaker3"],
categories_weight=[0.5, 0.3, 0.2]) # Proportions of annotations per speaker
reference_continuum: Continuum = sampler.sample_from_continuum
You could also measure the behavior of the gamma with your dissimilarity by tweaking the values in the sampler. Now, let’s use the corpus shuffling tool to generate a continuum with several annotators, with the selected errors with a given magnitude \(m\):
cst = CorpusShufflingTool(magnitude=m, # m is a float in [0, 1]
reference_continuum=reference_continuum)
generated_continuum: Continuum = cst.corpus_shuffle(
["Annotator1", "Annotator2", "Annotator3"],
shift=True, # annotations are randomly translated proportionally to m
false_pos=True, # random annotations are added, amount propotional to m
false_neg=True, # random annotations are discarded, amount propotional to m
split=True, # segments are splitted in two, number of splits propotional to m
cat_shuffle=True, # annotation categories are changed, amount propotional to m
include_ref=False) # If true, copies the reference's annotations.
Now you have a beautiful continuum, ready to be worked on !
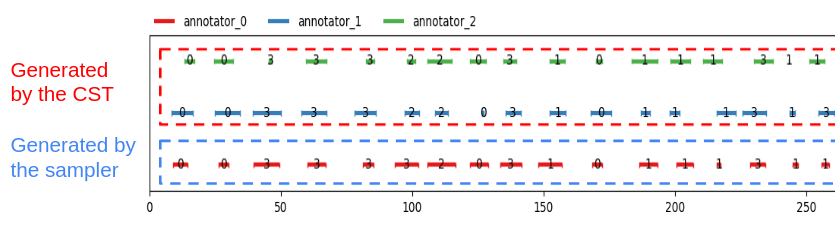
dissim: AbstractDissimilarity
gamma_results = generated_continuum.compute_gamma(dissim)
Beware that a lot of randomness is involved in gamma computation and continuum generation, so you might want to seed
using np.seed if you’re making graphs. Averaging several values computed from continua generated with the same
parameters might be better too.